With our January 2021 release, we've continued to add new tools and features to the Common Metabolic Diseases Knowledge Portal (CMDKP) and its component disease-specific portals.
See a summary of effector gene predictions
The CMDKP displays lists of predicted type 2 diabetes (T2D) effector genes generated by three different methods. In an effort to make it easier to compare their results, we've created a new interface, the T2D Effector Prediction Summary, that integrates the three sets of results. It’s available under the "Tools" menu of the CMDKP, T1DKP, and T2DKP.
On the summary interface, click on the "View Research Method" tab to see details on how the comparison between the three lists was made. In particular, note that the Integrated Classifier prediction method excludes the genes in the Curated T2D effector gene predictions list, since they were used as a gold-standard training set in developing that method.
At present, the effector gene lists are static, and two of them include predictions for T2D only. However, we're working to encode several methods for effector gene prediction in computational pipelines so that we can produce these lists for multiple phenotypes and update them regularly. This summary is a first attempt at integrating the results of the different effector gene prediction methods, and we welcome your feedback on how to improve it.
Stop by and play in the Knowledge Portal Labs
With a nod to Google Labs, described by Google as "a playground where our more adventurous users can play around with prototypes of some of our wild and crazy ideas and offer feedback directly to the engineers who developed them", we've created the Knowledge Portal (KP) Labs to showcase our in-progress work on new tools and interfaces. These are listed and described on the About KP Labs page.
Currently, KP Labs includes three tools:
- The Complications Association Browser allows you to compare lists of genes that are associated with a complication of T2D, such as peripheral vascular disease, with genes that are associated with that condition outside of the context of T2D.
- An all-new version of our classic Genetic Association Interactive Tool (GAIT) draws on exome sequence data from AMP T2D-GENES and TOPMed and offers 4 different aggregation test methods to facilitate custom gene burden tests for T2D and 24 quantitative phenotypes.
- The T2D Genetic Loci Clustering interface allows you to explore the results of a study that classified type 2 diabetes-associated genetic loci into 5 clusters by their associations with T2D-related traits, revealing possible disease mechanistic pathways (Udler et al. 2018).
The KP Labs pages are available under a new upper menu item on the CMDKP, T1DKP, and T2DKP. Please visit the Labs pages, kick the tires on the new tools, and let us know your thoughts on how to develop them further.
See new data and links in LocusZoom
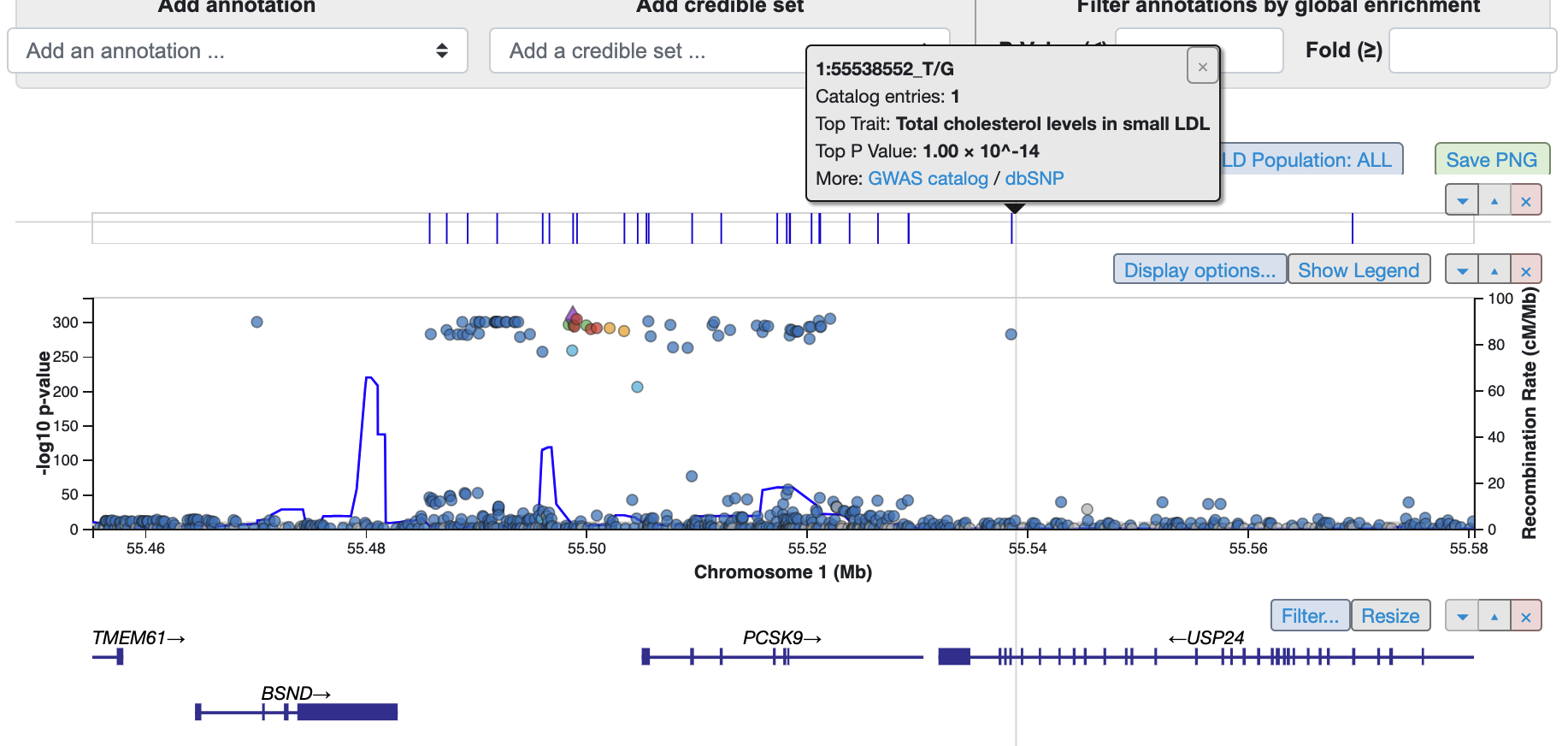
The LocusZoom interactive visualization within the GEnomic region Miner (GEM) module on the Region page has two new features. A track above the associations plot shows the genomic positions where variant associations are present in the GWAS Catalog. Mouse over a tick mark on the track to see more information and link to GWAS Catalog information for that position.
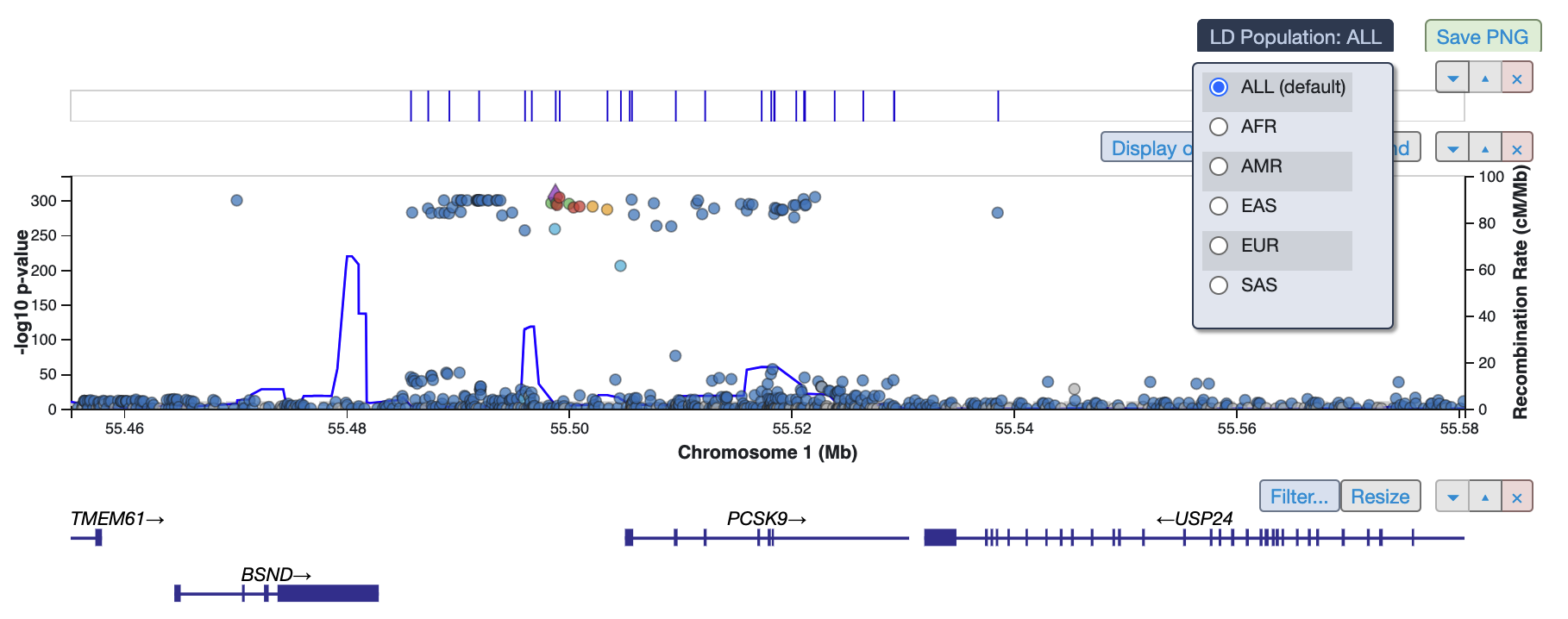
You can now also adjust the plot by choosing a specific linkage disequilibrium (LD) population:
Please try the new features and send us your feedback!
